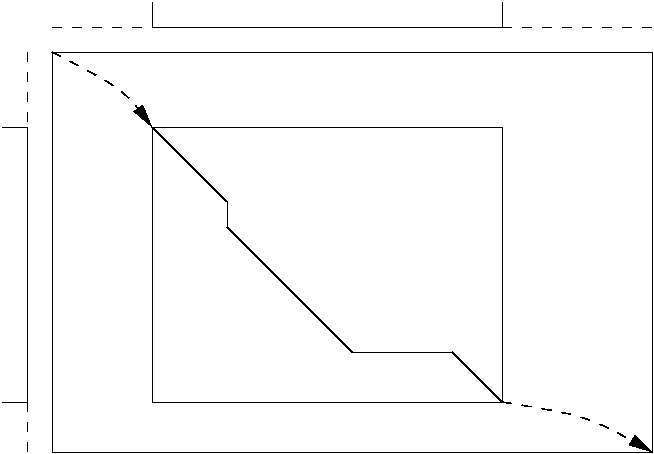
Local v. global.
A seminar to the Bioinformatics Group &
Department of Computer Science,
University of Wales Aberystwyth,

e.g.
Characters (bases, amino acids) are not all equal in all contexts, hence model in context (red):
|
|
r e a d o f f l i n e |
| ||
| generic (global, 1-state) DPA | (global, optimal) M-alignment |
|---|
Generalizes to k-states, local & global, optimal & summed.
ROC curves often used in this area.
Want high coverage (few false -ves) and
few errors (few false +ves),
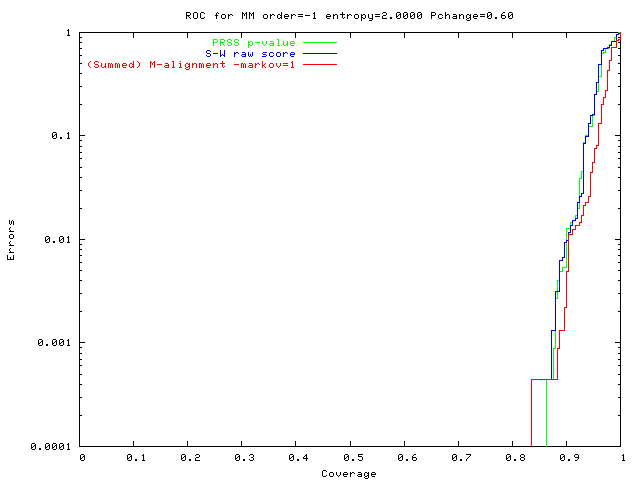 Easiest problem, new method ~ shuffling.
| |
|
green: PRSS p-value
(blue: S-W raw score); red: (summed) M-align (Markov=1). |
Uniform random pop'n (2 bits/base): All methods should do well, & they do. |
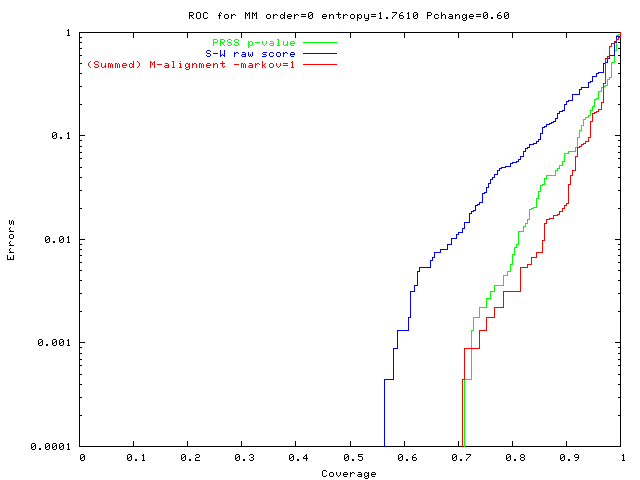 Easy problem, new method ~ shuffling
| |
|
green: PRSS p-value
(blue: S-W raw score); |
0-order data, biased composition: PRSS good, M-align best. |
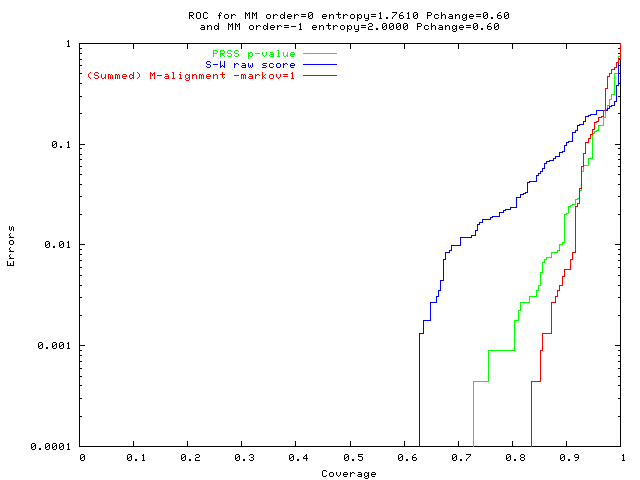 Harder problem, new M-alignment method (red) best
| |
|
green: PRSS p-value
(blue: S-W raw score); red: (summed) M-align (Markov=1). | Mixed pop'n, high entropy 0-order seq's and low entropy 0-order seq's |
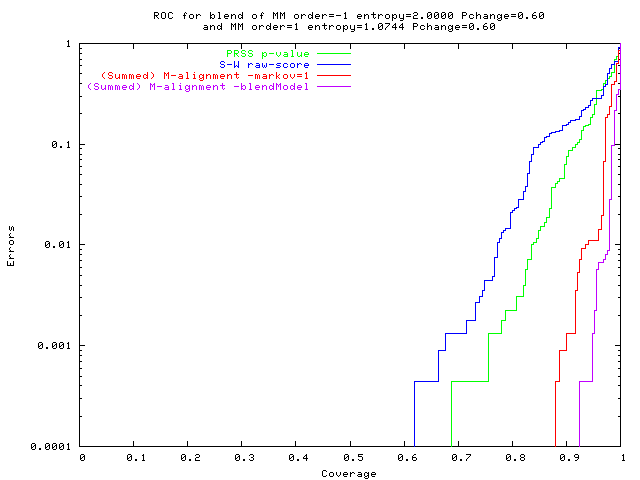 Hard problem,
new M-alignment method (red & purple) best
| |
|
green: PRSS p-value;
(blue: S-W raw score); red: (summed) M-align (Markov=1); purple: M-align (blended seq' model). | Pop'n of mixed seq's of high (2-bit/base) & low entropy 1st-order regions |
Reading:
[*] Like
many things out of Computer Science, Monash, this work owes a debt to